8.6 Cyclic Pattern Kernels
Cyclic pattern graph kernels [31,32] are based on the idea of mapping graphs to sets of cyclic and tree pattern strings that are compared using the intersection kernel.
8.6.1 Definition
A subgraph is a simple cycle if it is connected and each vertex has degree 2. Let S(G) denote the set of simple cycles in a graph G. An edge not belonging to a simple cycle is a bridge. We denote the subgraph of all bridges in G, which is a forest, by ![]() (Fig. 8.4). Let π be a mapping, computable in polynomial time, from the set of labeled simple cycles and trees to label strings that is injective modulo isomorphism. Note that such a mapping can always be constructed [31], [72]. The sets of cyclic and tree patterns are given by
(Fig. 8.4). Let π be a mapping, computable in polynomial time, from the set of labeled simple cycles and trees to label strings that is injective modulo isomorphism. Note that such a mapping can always be constructed [31], [72]. The sets of cyclic and tree patterns are given by ![]() and
and ![]() . The cyclic pattern kernel is given by
. The cyclic pattern kernel is given by
where ![]() denotes the intersection kernel.
denotes the intersection kernel.
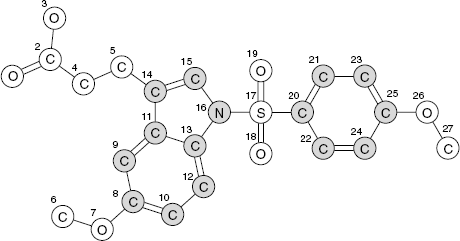
Figure 8.4 Cyclic and tree patterns of the molecular structure graph of indeglitazar. Shown are vertices belonging to simple cycles (shaded) and to bridges (white).
8.6.2 Computation
Computing Equation 8.18 is at least as hard as counting simple cycles in a graph, which is computationally not tractable if P ≠ NP [73]. For graphs with a small number of simple cycles, Equation 8.18 can be computed via enumeration of ![]() [31]. The number of cyclic and tree patterns in a graph can be exponential in |V|, leading to computational infeasibility of the cyclic pattern kernel for general graphs [31]. The restriction of inputs to graphs with few simple cycles can be relaxed to graphs of bounded treewidth, for which many NP-complete problems become tractable [74]. For graphs of constant bounded treewidth, Equation 8.18 can be computed in time polynomial in
[31]. The number of cyclic and tree patterns in a graph can be exponential in |V|, leading to computational infeasibility of the cyclic pattern kernel for general graphs [31]. The restriction of inputs to graphs with few simple cycles can be relaxed to graphs of bounded treewidth, for which many NP-complete problems become tractable [74]. For graphs of constant bounded treewidth, Equation 8.18 can be computed in time polynomial in ![]() [32].
[32].
8.6.3 Variants and Extensions
An alternative relaxation is to consider a different class of cycles. Horváth [32] uses algebraic graph theory to compute the cyclic pattern kernel on monotone increasing subsets of simple cycles generated by relevant cycles [75], with a similar runtime bound, but with different cyclic patterns. The number of relevant cycles is exponential in the worst case; for molecular graphs, it is typically cubic in |V| [76].
