Start a new R notebook and call it R Basics. We can enter a small script just so we can see how the steps progress for an R script. Enter the following into separate cells of your notebook:
myString <- "Hello, World!" print (myString)
You will end up with a starting screen that looks like this:

We should note the aspects of the R notebook view:
- We have the R logo in the upper-right corner. You will see this logo running in other R installations.
- There is also the peculiar R O just below the R icon. The unfilled circle indicates that the kernel is at rest, and the filled circle indicates the kernel is working.
- The rest of the menu items are the same as we have seen before.
This is a very simple script-set a variable in one cell then print out its value in another cell. Once executed (Cell | Run All), you will see your results:
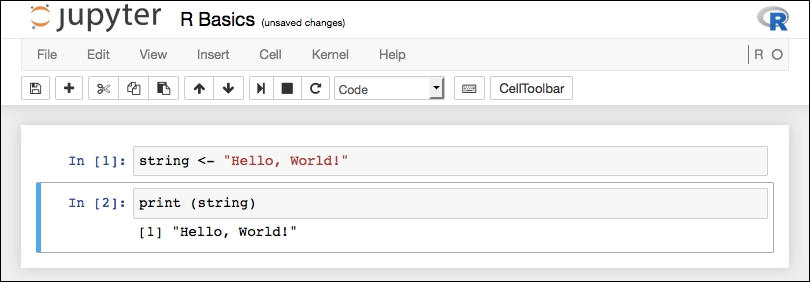
So, just as if you ran the script in an R interpreter, you get your output (with the numerical prefix). Jupyter has counted the statements so we have incremental numbering of the cells. Jupyter has not done anything special to print out variables for debugging, you would have to do that separately.
If we look at the R server-logging statements (a command-line window was created when we started Jupyter), we see the actions that took place:
$ jupyter notebook [I 11:00:06.965 NotebookApp] Serving notebooks from local directory: /Users/dtoomey/miniconda3/bin [I 11:00:06.965 NotebookApp] 0 active kernels [I 11:00:06.965 NotebookApp] The Jupyter Notebook is running at: http://localhost:8888/ [I 11:00:06.965 NotebookApp] Use Control-C to stop this server and shut down all kernels (twice to skip confirmation). [I 11:00:17.447 NotebookApp] Creating new notebook in [I 11:00:18.199 NotebookApp] Kernel started: 518308da-460a-4eb9-9959-1411e31dec69 [1] "Got unhandled msg_type:" "comm_open" [I 11:02:18.160 NotebookApp] Saving file at /Untitled.ipynb [I 11:08:27.340 NotebookApp] Saving file at /R Basics.ipynb [1] "Got unhandled msg_type:" "comm_open" [I 11:14:45.204 NotebookApp] Saving file at /R Basics.ipynb
We started the server, created a new notebook, and saved it as R Basics. If we open the IPYNB file on disk (using a text editor), we can see the following:
{
"cells": [
...<similar to previously displayed>
],
"metadata": {
"kernelspec": {
"display_name": "R",
"language": "R",
"name": "ir"
},
"language_info": {
"codemirror_mode": "r",
"file_extension": ".r",
"mimetype": "text/x-r-source",
"name": "R",
"pygments_lexer": "r",
"version": "3.2.2"
}
},
...<omitted>
}
This is a little different than what we saw in the prior chapter on Python notebook coding. Particularly, the metadata clearly tells the script cells to be R script. Note, the actual cells are not specific to a language-they are just scripts that will be executed as per the metadata directives.
