We can use map/reduce to estimate the Pi. Suppose we have code like this:
import pyspark import random if not 'sc' in globals(): sc = pyspark.SparkContext() NUM_SAMPLES = 1000 def sample(p): x,y = random.random(),random.random() return 1 if x*x + y*y < 1 else 0 count = sc.parallelize(xrange(0, NUM_SAMPLES)) .map(sample) .reduce(lambda a, b: a + b) print "Pi is roughly %f" % (4.0 * count / NUM_SAMPLES)
This code has the same preamble. We are using the random Python package. There is a constant for the number of samples to attempt.
We are building an RDD called count. We call upon the parallelize function to split up this process over the nodes available. The code just maps the result of the sample function call. Finally, we reduce the generated map set by adding all the samples.
The sample function gets two random numbers and returns a 1 or a 0 depending on where the two numbers end up in size. We are looking for random numbers in a small range and then comparing whether they occur within a circle of the same diameter. With a large enough sample, we would end up with Pi (3.141...).
If we run this in Jupyter, we see the following:
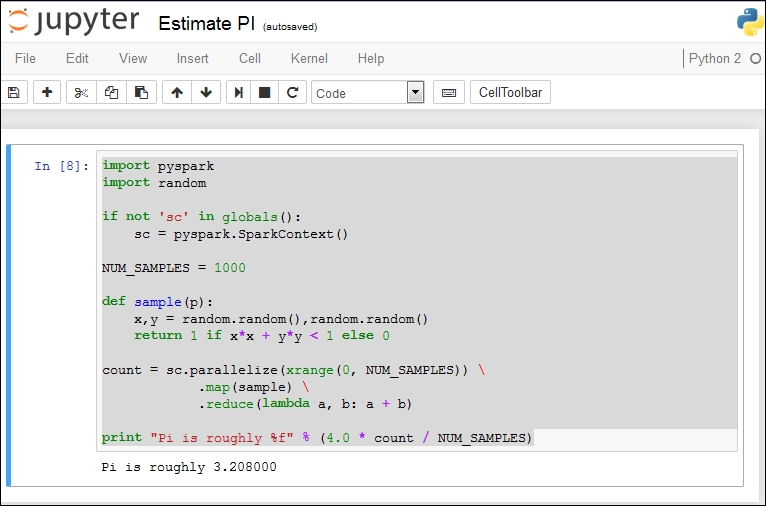
When I ran this with NUM_SAMPLES = 10000, I ended up with this:
PI = 3.138000.
