5. An entire industry from a single cell
The biotechnology industry arrived in the late 1970s when entrepreneur-biologists began harnessing microbes for profit. A new company, Genentech, first entered the commercial market in 1977 with the peptide somatostatin, made by E. coli engineered to carry genes that encoded for this growth-modulating hormone. Prior to these E. coli fermentations, somatostatin came only from cattle after slaughter.
The first success in moving genes from one organism into a different, unrelated organism occurred in 1972 in Paul Berg’s laboratory at Stanford University. Berg composed a hybrid DNA from the DNA molecules extracted from two different viruses. The next year Herbert Boyer and Stanley Cohen further stretched the boundaries of gene transfer by putting genes from a toad into E. coli. Most important, successive generations of the engineered E. coli retained the new gene and reproduced it whenever they made new copies of E. coli genes. Boyer and Cohen had developed recombinant DNA, and as a result, the world had its first human-made GMO.
Some biotechnologists have taken a broad view of when their science began, citing the first use of bacteria or yeasts to benefit humans. Using this criterion, biotech began in 6000 BCE when people first brewed beverages using yeast fermentations. For practical purposes, the science of manipulating microbial, plant, and animal genes emerged when scientists first cleaved DNA and then inserted a gene from an unrelated organism into it. The biotech industry commenced when companies made the first commercial products from recombinant DNA by growing large volumes of GMOs.
Berg, Boyer, and Cohen would not have initiated the new science of genetic engineering without prior individual accomplishments in genetics. Walther Flemming, in 1869, collected a sticky substance from eukaryotic cells he called chromatin, later to be identified along with associated proteins as the chromosome. In most bacteria, the chromosome is a single DNA molecule packed into a dense area of the cell (called DNA packing). Bacteria do not contain the proteins, called histones, which eukaryotes use for keeping the large DNA molecule organized. Eukaryotes carry from one to several chromosomes. The eukaryotic chromosomes plus DNA located in mitochondria collectively make up the organism’s genome. In bacteria, the genome consists of the DNA plus plasmids.
In the early 1900s, Columbia University geneticist Thomas Hunt Morgan used Drosophila fruit flies to demonstrate that the chromosome, in other words DNA, carried an organism’s genes. Less than 50 years later, American James Watson and British Francis Crick, who were both molecular biologists, described the structure of the DNA molecule.
DNA structure resembles a ladder that has been twisted into a helix. The long backbones, or strands, consist of the sugar deoxyribose, each holding a phosphate group (one phosphorus connected to four oxygens) that extends away from the ladder. Deoxyribose also holds a nitrogen-containing base on the opposite side that holds the phosphate group. Each base points inward so that different bases from each strand and complementary in structure connect by a chemical bond. These bonds, called hydrogen bonds, hold atoms together by weak connections compared with other types of chemical bonds.
Nature uses only four bases in DNA to serve as a type of alphabet. These bases are adenine, thymine, cytosine, and guanine, which biologists abbreviate to A, T, C, and G, respectively. The sequence of bases in DNA determines the makeup of genes, which are short segments of bases. The exact sequences of A, T, C, or G in each living organism hold all of the genetic information that defines the organism’s species and also makes every individual unique. No two DNA compositions are identical.
Paul Berg and the other leading molecular biologists first created hybrid DNA by cutting the strands with an enzyme called restriction endonuclease. (Restriction endonucleases evolved in bacteria for the purpose of destroying foreign DNA brought into the cell by an invading phage.) The break in the DNA molecule served as a place to insert one or more genes from another organism.
An alphabet composed of only four letters does not seem adequate for carrying all the heredity of every organism on Earth. Nature solved this potential problem by requiring that each sequence of three bases serve as the main unit of genetic information, called the genetic code. The base triplet constitutes a codon, and each codon translates to one of nature’s amino acids, which act as the building blocks of all proteins regardless of whether the proteins belong to animals, plants, or microbes. Only 20 different amino acids go into nature’s proteins that vary in length from about 100 to more than 10,000 amino acids. The three-letter codons increase nature’s capacity to put all of its information into genes made of no more than four letters. The varying lengths of proteins further expand the possibilities for defining everything in nature from a simple microbe to a human. The genetic code furthermore defines every being that once lived but has gone extinct.
Imagine if only one base were to encode for one amino acid. Proteins would not be able to contain more than four amino acids. A codon composed of two bases could hold a maximum number of amino acids of 42,, or16. By adding one more base, the maximum number of amino acids that the alphabet could define would be 43, equal to 64. DNA’s triplet codons can thus identify all of the essential amino acids with several codons to spare. Because nature tries to do things in the simplest way possible, it has no need to design four-, five-, or longer base codons to accomplish the same job performed buy a three-base codon.
Nature makes use of the extra 44 codons that do not translate directly to an amino acid by assigning some of them specific meanings, such as “The gene starts here” and “The gene stops here.” The genetic code, unlike the 26-letter alphabet used in English, contains redundancy but no ambiguity. Redundancy allows some amino acids to have more than one codon that defines them. For example, DNA uses either of two codons to spell the amino acid arginine (AGA and AGG), but six different codons can each spell the amino acid serine. No ambiguity occurs in the genetic code, however, because no codon ever specifies more than one amino acid. Contrast the genetic code to the English alphabet containing the five-letter heteronym “spring,” which could mean a mechanical device inside a mattress, a freshwater source, the act of leaping, or a season.
Redundancy helps biological systems operate with some versatility so that even a slight mistake in a base sequence can translate into the correct amino acid used for building a protein. Cells also contain repair systems that proofread the code. Repair system enzymes excise incorrect bases, fix mismatched bases in the ladder’s rungs, and rebuild damaged sections of DNA.
The genetic code connects all biological organisms. Regardless of the organism from single-celled bacteria to the most complex—usually assumed by egocentric humans to be the human—all use the same genetic alphabet to define amino acids and thus proteins. The universal nature of the genetic code allows scientists to study E. coli for the purpose of learning about human genes. In addition, the unity of biology makes the opportunities for genetic engineering almost unlimited because every organism uses the same basic means of building its cellular constituents.
Genetic engineering has not replaced the chemical industry, although industry leaders in Europe have made plans to convert chemical manufacturing processes to biological processes. This new business model, called white biotechnology, uses bacteria or their enzymes to carry out manufacturing steps that presently require high heat and hazardous catalysts. White biotech produces no hazardous waste and requires much less energy input than conventional manufacturing. The U.S. biotech industry familiar to most people and responsible for making GMOs is called green biotechnology. The biotechnology industry currently designates color codes to specific areas of interest:
• Green—Bioengineered microbes, food crops, and trees
• White—Microbial enzymes applied to industrial manufacturing
• Blue—Biotechniques oriented toward marine biology
• Orange—Engineered yeasts
• Red—Medical gene therapy, tissue therapy, and stem cell applications
In the 1950s, companies rebuilt their businesses for a peacetime economy. The chemical industry had been expanding since the 1930s and boomed in the 1950s with a new mantra: convenience products. DuPont Company communicated its industry’s bright future as well as its customers’ with the slogan “Better Things for Better Living... Through Chemistry.” By 1964’s New York World’s Fair, DuPont had rolled out a song-and-dance extravaganza on the power of chemistry. The industry’s new drugs, pesticides, and plastics promised a better quality of life, but these products also required significant quality control at the manufacturing level. Wartime expertise in physics and chemistry turned toward making new analytical equipment to inspect a compound’s structure and measure its purity. Companies such as Hewlett-Packard, Varian Associates, and Perkin-Elmer filled the gap.
Alec Fleming’s legacy inspired a new interest in biology in the 1940s, but additional breakthroughs came slower than many people might have expected. Antibiotic discovery involved laborious manual tests. Microbiologists scooped up soil samples, recovered the soil’s fungi and bacteria, and then searched for extracts from the cultures to test against hundreds of bacteria. In addition to the tedium, microbiologists often saw variable results in laboratory tests. When a microbiologist inoculates ten tubes of broth with the same Staphylococcus, eight tubes might grow, one tube might not grow, and the tenth tube ends up contaminated. Chemists at drug companies sped up the process by synthesizing new antibiotics based on the structures of known natural antibiotics. By the 1950s, the chemical industry offered a faster way to find new drugs. To keep up with the chemists, microbiologists needed a dependable microbe that grew easily and quickly to large quantities.
E. coli
In the 1880s, outbreaks of infant diarrhea raged through European cities and killed hundreds of babies. Like other physicians, Austrian pediatrician Theodore von Escherich struggled to save his patients and simultaneously find the infection’s cause. He recovered various bacteria from stool samples without an idea as to their role, if any, in the illness. In 1885 von Escherich published a medical article describing 19 bacteria that dominated the infants’ digestive tracts. One in particular seemed to be consistently present and in high numbers. He named it (with a striking lack of creativity) Bacterium coli commune for “common colon bacterium.” In 1958, the microbe was renamed Escherichia coli in honor of its discoverer.
E. coli’s physiology offers nothing remarkable. It does not pump out an excess of useful or unique enzymes or make antibiotics. It dominates the newborn’s intestinal tract but gradually other bacteria overtake it and carry out the important microbial reactions of digestion. For example, strict anaerobes produce copious amounts of digestive enzymes that help break down proteins, fats, and carbohydrates. These bacteria also partially digest fibers and synthesize proteins and vitamins that are used in the host’s metabolism. E. coli does not contribute as much to digestive activities as the strict anaerobes, but because it is a facultative anaerobe that uses oxygen when present and lives without oxygen in anaerobic places, its main role is to deplete oxygen so that anaerobic bacteria can flourish.
Von Escherich probably noticed that E. coli grows fast to high numbers in laboratory cultures. The species flourishes on a wide variety of nutrients and does not need incubation. Leave a flask of E. coli on a lab bench overnight and a dense culture will greet the microbiologist the next morning. The strict anaerobes of the digestive tract take three days or longer to grow to densities that E. coli reaches in about 10 hours.
By the turn of the century, doctors had not solved the problem of infant diarrhea—it remains a significant worldwide cause of infant mortality. They did, however, think that E. coli might be useful for treating intestinal ailments in adults. In Freiburg, Germany, physician Alfred Nissle planned to use E. coli for intestinal upsets such as diarrhea, abdominal cramping, and nausea. In this so-called “bacteria therapy” Nissle believed that dosing an ill person with live E. coli might drive the pathogenic microbes from the gut.
From 1915 to 1917, Nissle tested various mixtures of E. coli strains in Petri dishes against typhus-causing Salmonella. When a mixture appeared to be antagonistic toward the Salmonella, he tried it on other pathogens. Nissle finally concocted a “cocktail” of what he considered the strongest E. coli strains and with considerable courage he drank it. When no harmful effects ensued, Nissle felt he was on the road to an important medical discovery.
During the period Nissle had been conducting his E. coli experiments, Germany’s army suffered from severe dysentery as had others throughout Europe during the Great War. Dirty water, bad food, and exhaustion conspired to weaken men in the foxholes as well as civilians. In 1917 Nissle made his way to two field hospitals in search of a super E. coli that would work even better than the strains in his laboratory. In one tent, he found a non-commissioned officer who had suffered various injuries but never fell victim to diarrhea even when everyone around him had it. Nissle cultured some E. coli from the soldier and returned to Freiburg.
Alfred Nissle grew the special E. coli in flasks and then poured it into gelatin capsules. When the job of supplying an entire army overwhelmed him, he commissioned the production to a company in Danzig. The new antidiarrheal capsules were called Mutaflor. The wartime upheavals in Europe through 1945 forced Nissle to move the manufacturing more than once, but the production of Mutaflor never ceased. Mutaflor remains commercially available today as a probiotic treatment for digestive upset. The product contains “E. coli Nissle 1917” made from direct clones of the superbug Nissle isolated on the battlefield in 1917. The original strain that Nissle submitted to the German Collection of Microorganisms remains in its depository in Braunschweig.
At Stanford University in 1922, microbiologists noted another fast-growing E. coli strain with a curious trait: It did not cause illness in humans. The strain received a laboratory identification of “K-12.” K-12 became a standard in teaching and research laboratories, soon shared by Stanford with other universities. When eventual Nobel Prize awardees Joshua Lederberg and Edward Tatum began studies on how genes carry information and the mechanisms organisms use to exchange this information, they made the logical choice of K-12 as an experimental workhorse. E. coli became forever linked with advances in genetics and biotechnology.
Since the first K-12 experiments, more than 3,000 different mutants of this bacterium have been used in cell metabolism, physiology, and gene studies. One of the first bacterial genomes to be sequenced was that of K-12; the complete sequence of its 4,377 genes was published in 1997. In the last 50 years, 14 Nobel Prizes have been awarded based on work done with E. coli, mainly K-12.
The power of cloning
By the 1970s, microbiologists were routinely taking apart E. coli to study its reproduction, enzymes, and virulence. The chemical industry had lost some luster with each new discovery of environmental pollution, and biology again looked like the science for the future: clean, quiet, and nonpolluting.
In biotech’s infancy, “cloning” became the buzzword for the power of this new technology: The ability to take a single gene and produce millions of identical copies. E. coli became a living staging area in which genes were cloned by the following general scheme:
- Extract DNA from an organism possessing a desirable trait (a gene).
- Cleave the DNA into many smaller pieces with a specialized enzyme called restriction endonuclease (RE).
- Extract E. coli plasmids and open their circular structure with another RE.
- Insert the various DNA fragments into the many plasmids.
- Allow the bacteria to take the plasmid back into their cells.
- Grow all the bacteria and use screening procedures to identify the cells carrying the desirable gene.
- Grow large amounts of these gene-carrying cells, that is, cloning.
- Harvest the product that the gene controls.
In biotech’s infancy scientists painstakingly worked out each of the preceding steps (see Figure 5.1). Molecular biologists perfected the art of extracting DNA from cells without breaking the large molecule into pieces. They devised techniques for splicing new segments of one type of DNA into a second DNA molecule, and they developed methods for testing the activity in a new GMO. But the scientists also noticed that their favorite bacterium E. coli resisted taking plasmids into their cells, a key step in genetic engineering called transformation. Without an easy way to deliver DNA into E. coli, many genetic experiments might become impossible. In 1970, Morton Mandel and Akiko Higa solved this dilemma by showing that calcium increased the permeability of cell membranes to DNA. By soaking E. coli in a chilled calcium chloride solution for 24 hours, biologists now make the bacterium 20 to 30 times more receptive to taking up plasmids. Bacteria that take in plasmids from the environment are called competent cells, and biotechnologists now use this simple soaking step to make E. coli competent for transformation.
Figure 5.1. Microbiologists in environmental study, medicine, industry, and academia use the same aseptic techniques. These disciplines use methods adapted from biotechnology for manipulating the genetic makeup of bacteria.
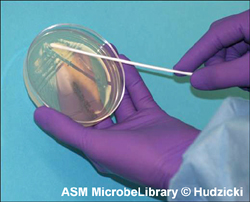
(Reproduced with permission of the American Society for Microbiology MicrobeLibrary (http://www.microbelibrary.org))
In the early days of biotech research, bacterial cloning—it used to be called gene splicing—served as the only way to make large amounts of gene and gene products. Bacteria make millions of copies of a target gene by replicating it each time a cell splits down the middle to make two new cells, a process called binary fission. In time, scientists developed methods for using bacteriophages to deliver genes directly into a bacterium’s DNA. Viruses’ modus operandi involves appropriating a cell’s DNA replication system, which is a perfect mechanism for delivering a foreign gene into bacterial DNA. PCR would enter the picture next as a faster DNA-amplification method.
E. coli remains a major tool in biotechnology, but additional microbes such as the yeast Saccharomyces cerevisiae and the bacterium Bacillus subtilis also contribute a large share to recombinant DNA technology. Biotech companies use the basic cloning scheme described previously in yeasts and bacteria for making the drugs listed in Table 5.1.
Table 5.1. Major products made by biotechnology
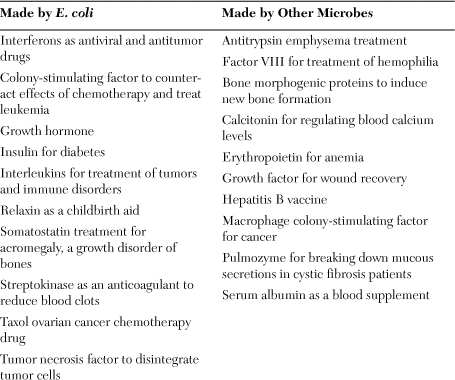
Biotech companies manufacture their products in fermentation vessels of 300 to 3,000 gallons. Technicians at biotech companies scale up bacterial cultures from small volumes of less than a gallon to fermenters of several gallons. After this modest scaling up of the culturing process, workers in a manufacturing plant increase the production size more by growing the GMO in vessels of 300 to 3,000 gallons. All of the actions leading up to large-scale production comprise upstream processing. A different team of technicians monitors downstream processing, which encompasses all the steps from fermentation to the packaging of a clean, pure final product. Genentech and Amgen, both in California, became the first two biotech companies to reach this large-scale level of production.
In 1996, scientists at Scotland’s Roslin Institute created Dolly, the first mammal (a sheep) made by cloning DNA from an adult animal. The public and many scientists reacted to this news with concern that humans would be the next organism to be cloned. Renee Reijo Pera, stem cell researcher at the University of California-San Francisco, remarked, “You can almost divide science into two segments: Before Dolly and After Dolly.” But cloning higher organisms had little in common with bacterial cloning for making GMOs. Dolly’s clone came about by transferring the nucleus—containing an animal’s entire genome—from an adult sheep’s cell into mammary tissue where the genome replicated as the tissue reproduced. Cows, goats, pigs, rats, mice, cats, dogs, horses, and mules have since been similarly cloned.
The goal of animal cloning is to produce a new animal identical in every way possible to the original animal. Animal cloning thus seeks to repeat an entire genome in a new animal. Gene cloning in bacteria, by contrast, serves as a simple way to make many copies of one or more genes in a short period of time. In short, animal cloning makes new animal copies, and bacterial cloning makes new gene copies. By inserting one or more genes into a bacterial cell’s DNA and then growing the cells through several generations, a microbiologist can produce millions of copies of the “new” DNA overnight because of bacteria’s fast growth rate.
A chain reaction
One spring evening in 1983, biochemist Kary Mullis drove from his job at Cetus Corporation near San Francisco to his cabin in California’s quiet Anderson Valley. The San Francisco Bay Area had just begun to plant the seeds for the new science called biotechnology. Molecular biologists had learned how to open DNA molecules with enzymes and insert genes from an unrelated organism. But cloning bacteria to make each new batch of genes required considerable labor, and the bacterial cultures produced only miniscule amounts of desired DNA. Mullis pondered this problem as he drove Route 128. He recalled reading of a bacterium living in hot springs and containing enzymes active at high temperatures that melted most other enzymes. Before reaching his cabin, Kary Mullis had developed an idea that would revolutionize biology.
In 1966, famed microbial ecologist Thomas Brock and graduate assistant Hudson Freeze had discovered a bacterium surviving the blistering conditions of Mushroom Spring in Yellowstone National Park. They named the species Thermus aquaticus, Taq for short, and sent a culture to a national repository for microbes near Washington. DC. Various microbiologists studied the thermophile and its enzymes, but Taq seemed to offer little in the way of useful attributes. Mullis suspected that Taq in fact held a crucial attribute.
At a temperature approaching 200°F, DNA becomes unstable and separates, or denatures, into two single strands instead of its normal double stranded confirmation. Back in his lab, Mullis raised the temperature of a DNA mixture to denature the molecule and then added DNA fragments called primers plus the enzyme DNA polymerase he had extracted from Taq. Next, Mullis lowered the temperature to about 154°F wherein the polymerase began building new DNA copies from the old strands and the primers. By repeatedly heating and cooling the mixture, Mullis could produce about one million copies of the new DNA in 20 minutes and a billion copies in 30 minutes. Molecular biologists call this production of millions of DNA copies from a small, single piece of DNA, amplification. Taq’s DNA polymerase provided the key to Mullis’s invention because it withstands repeated heating to very high temperatures and then carries out the DNA synthesis step at the cooler (but still high) temperature.
The new process called polymerase chain reaction (PCR) made any snippet of microbial DNA analyzable. Michael Crichton capitalized on PCR’s extraordinary potential in the 1990 book Jurassic Park, in which scientists amplify dinosaur DNA preserved in ancient amber. Although PCR can amplify pieces of DNA that had been dormant in nature for years, Jurassic Park’s re-creation of an entire extinct genome seemed implausible when the movie was released because closing missing gaps was error-prone. Today, computer programs calculate the likely base sequences of missing pieces of DNA to fill in gaps in damaged DNA. As the power of these programs increases, scientists will reconstruct extinct DNA with increasing accuracy.
When a character on a television crime show says, “We need a rush on the DNA,” a harried lab technician produces within minutes the name (accompanied by an up-to-date photo) of the bad guy on a computer screen. These scenes depict the power of PCR for analyzing biological matter, but the entire PCR process actually takes much longer. A technician first prepares the DNA-primer-polymerase mixture. A heating-cooling machine called a thermocycler requires at least 2 hours to amplify the bits of DNA. Next, the scientist must determine the sequence of the DNA’s subunits or bases, which takes another 24 hours if using an automatic sequencer instrument. Lacking such an machine, manual methods can take up to 3 weeks.
The Food and Drug Administration (FDA) and other government agencies have put PCR to use in crime-solving. In early 2009, the FDA began a food product recall that would encompass 3,900 peanut butter products as suspect causes of a nationwide Salmonella outbreak that sickened 700 people and killed nine. Microbiologists used PCR to amplify the DNA from bacteria in the products and determine the pathogen’s unique sequence. This so-called DNA fingerprint led CDC investigators to a Blakely, Georgia, manufacturing plant. A leaky roof had allowed rain contaminated with Salmonella-laced bird droppings to land directly on food processing equipment and perhaps directly into peanut butter paste as well. Microbiologists can now trace a single pathogen strain from a person’s stool sample to an individual farm, a certain shift on the packing line, and even a specific agricultural field.
Real-time PCR has come on the scene as a faster way to analyze samples to prevent crimes from growing cold. In real-time PCR, a detector monitors the formation of increasing amounts of DNA in the thermocycler as it occurs, unlike traditional PCR that takes extra days to analyze the final products from the thermocycler step. Real-time RCR has helped fight the global poaching industry that trades hides, pelts, internal organs, horns, feathers, and shells of endangered animals, as well as caviar. Microscopic drops of an animal’s blood on a suspected poacher’s clothes can solve a case. Analysis of ivory sold on the black market has traced the ivory to specific elephant herds in Africa and sometimes to individual families.
Kary Mullis received the 1993 Nobel Prize in chemistry for developing PCR technology. Shortly afterward, the enigmatic Mullis joined a campaign to question the idea that HIV causes AIDS.
Bacteria on the street
Biotech has developed into a science of contradictions. For instance, few graduate students in microbiology complete their studies without doing some sort of gene sequencing or gene engineering. The majority of these students have no exposure at all to culturing whole bacterial cells except E. coli, and they spend more time with the disassembled bacterium than the whole living cell. The biotech business has a similar dichotomy. The earliest biotech advocates touted gene cloning as a step toward curing humanity’s worst diseases while antibiotech groups predicted the end of nature as we know it. Government leaders recognized the upside for the United States as a world leader in the emerging technology, but they also worried about the need to contain the frightening creatures about to emerge.
A conflicted Wall Street put a modicum of trust in the new industry but did not leap into the biotech pool with both feet. The public’s worry over safety did not make for attractive investments. Biotechnology’s proteins and cells rather than widgets also presented a new business model. What is a marketable product from a biotech company? Is it the cells that produce a hormone, the gene that encodes for the hormone, or the hormone itself? The U.S. Supreme Court helped clear up some of the confusion by ruling in 1980 that bioengineered bacteria could be patented.
At the start of the 1990s, biotech stocks rode the high-tech wave on Wall Street. By the mid-nineties, however, meager returns turned off the investors, and their interest in biotech cooled. Biotech-produced drugs were not easy to make. The manufacture of genetically altered organisms could produce surprises for even seasoned microbiologists. Mutant cells, contamination, and the capability of bacteria to shift their metabolic pathways slowed the early progress. Biotech’s biggest drawback resided in the fact that people could not make up their minds if the new technology was about to save them or kill them.
Warren Buffett described the perfect product, cigarettes, when he said, “It cost a penny to make. Sell it for a dollar. It’s addictive.” The dotcom industry grew based on this very philosophy. Biotechnology has not come near matching Buffett’s three criteria. Like conventional drugs, biotech products require large amounts of research cash and lengthy clinical testing on human subjects. Some drugs, such as new antibiotics, have become too expensive to develop. If people cannot imagine how they would exist without computer technology, they certainly can and do imagine a world without biotech products. In fact, an increasing number of people in the United States prefer a world without GMOs. Do we need GMOs? Bioengineered tomatoes taste fine, but so do organic, non-GMO varieties. Bioengineered bacteria clean up oil spills, but so do bacteria native to the waves and sands slickened with oil.
Biotechnology received a golden opportunity on March 24, 1989, to show the world the value of GMOs released into the environment for a purpose. The Exxon Valdez oil tanker hit a reef that day in Alaska’s Prince William Sound and spilled an estimated 11 million gallons of crude oil. Whipped by winds, about four million gallons of foamy crude washed ashore, coating 1,300 miles of coastal habitat for marine organisms, terrestrial animals, and birds. Marine bacteria at the spill burst into rapid growth in response to the influx of nutrients; crude oil provides a digestible carbon source for bacteria as opposed to refined oils. The United States did not permit the release of GMOs into the environment in an uncontrolled manner, so microbiologists could not put to work fast-growing bacteria engineered for oil degradation. They turned instead to bioaugmentation to carry out the largest microbe-based pollution cleanup project in history.
The Environmental Protection Agency’s John Skinner pointed out shortly after the spill, “Essentially, all the microorganisms needed to degrade the oil are already on the beaches.” Microbiologists accelerated the bacteria’s growth rate by adding nitrogen and phosphorus to the soils. Like bacteria fed a nutrient-rich broth in a laboratory test tube, the native bacteria responded to the augmentation of their environment with added nutrients. The bioaugmentation of the shore’s native bacteria (mainly Bacillus) is believed to have increased by at least sixfold the rate of oil decomposition.
GMOs fitted with the genes from native bacteria that degrade fuels, pesticides, industrial solvents, and toxic metal compounds have been developed in microbiology laboratories all over the world. Government agencies have slowed this progress by limiting GMOs to experimental study rather than real-life environmental disasters. The U.S. Environmental Protection Agency reserves the term “bioremediation” for pollution cleanup by unaltered native bacteria and not bioengineered strains. Communities may be nearing a point at which they must choose between hazardous substances in their environment and GMOs released to clean up those substances.
Biotechnology critics have warned of a future in which foods come from 3,000-gallon vats of bacteria. Economic and technology consultant Jeremy Rifkin has cautioned that bacteria will make soil and farms obsolete. I am not sure how this could ever happen. I suspect Rifkin is speaking in metaphor to warn us of GMO-produced foods that could supplant traditional agriculture. Rifkin’s Web site warns also that “the mass release of thousands of genetically engineered life forms into the environment [will] cause catastrophic genetic pollution and irreversible damage to the biosphere.” Biotechnology today must stand up to strong criticism even as it develops new life-saving drugs and invents processes that clean the environment.
The concept of developing microbial food for humans relates mainly to single-cell protein, or the use of microbial cells as a dietary protein supplement. This idea is at least 20 years old and had been proposed as a way of alleviating global hunger and protein deficiency. Single-cell protein from bacteria never reached practical levels for two reasons. First, bacteria grown in large quantities as a food must be cleared of any toxin or antibiotic they might make, which complicates the production process and raises costs. Second, microbial products packaged as a protein-dense food would likely induce serious allergic reactions in many consumers. Bacteria cannot replace traditional agriculture even if a future generation of scientists found a way to do it. The Earth needs green plants as much as it needs bacteria.
Warnings by biotech’s critics about GMOs invading natural ecosystems continue. Bacteria are survivors because of supreme adaptability. Could the adaptability needed by a GMO to carry out its job in nature also allow the microbe to take over ecosystems? Natural ecosystems possess exquisite mechanisms to ensure balance between competitive species from bacteria to higher organisms. Motility, quorum sensing, spore formation, and antibiotic production are examples of the many devices bacteria use to ensure they receive adequate habitat, nutrients, and water. A GMO must overcome all competitors to take over an ecosystem, but nature long ago developed mechanisms to protect the balance of species and resist drastic change. It is wise to remember also that bacteria have been exchanging genes that help them survive since the beginning of their existence. Most new genes that become part of a cell’s DNA by either gene transfer or mutation give no benefits to the cell. Because the genes of GMOs are designed to accomplish very specific tasks, the chances of a GMO ruling over natural communities seem remote.
The National Institutes of Health (NIH) have published hundreds of pages of regulations on GMOs with the intent of reducing the chances of a GMO accidentally escaping into the environment. These rules cover methods for containing recombinant microbes by physical, chemical, and biological tactics. Current physical containment involves the safe handling and disposal of GMOs so that live cells do not accidentally escape a laboratory and enter an ecosystem. Microbiologists use special safety cabinets based on the principles of BSL-4 cabinets. They also sterilize all wastes before discarding them. Chemical methods include disinfectants and radiation to kill bacteria in places they might have contaminated. But the dexterity with which bacteria evade chemicals—imagine the consequences of a GMO lodged inside a biofilm—highlights the weaknesses of chemical containment. To date, biological methods for making GMOs safe in the environment offer the greatest promise.
Microbiologists can engineer bacteria to self-destruct by adding suicide genes to recombinant DNA. Suicide genes control GMOs after the microbe has completed its task. The safety mechanism works by either positive or negative control. In both cases, a second compound, or activator, keeps the suicide gene from working until conditions change in the environment. In positive control, a chemical or other stimulus such as a certain temperature, affects the activator, which then releases its control over the suicide gene. The now active suicide gene initiates a progression of events in the cell that lead to its death, a process called apoptosis. In the hypothetical example already mentioned, a 3,000-gallon batch of bioengineered E. coli making growth hormone, ruptures and spills in a 7.0 Richter earthquake—a believable event in California where hundreds of biotech companies exist. The E. coli might rush into nearby soils and streams, producing hormone that traumatizes ecosystems. But a suicide gene designed to turn on when cells are exposed to a temperature of 72°F or lower—fermenters usually run at about 100°F—ensures that the E. coli destroys itself as soon as it escapes into the environment. Negative control turns on when a stimulus in the environment disappears. Bioremediation bacteria designed to degrade a pollutant, for example, undergo apoptosis when the pollutant disappears.
E. coli is the world’s most bioengineered microbe and also provides suicide genes for other GMOs. The gef gene in E. coli encodes for a 50-amino acid protein, small by protein standards, that turns on apoptosis in several different bacterial species. The gef gene has already been investigated as a treatment against melanoma cells and breast cancer, and for controlling engineered Pseudomonas. Bioengineered P. putida degrades alkyl benzoates, which are thickeners used in cosmetic products and drugs. As long as this pollutant remains in the environment, P. putida equipped with E. coli’s gef gene continues breaking it down. When the pollutant level has been reduced, the gef protein interferes with the normal flow of energy-producing electrons in P. putida’s membrane. The bioengineered Pseudomonas commits suicide.
Biological containment systems control GMO cells from within the cell and thus promise the best method of preventing GMO accidents. But one P. putida cell per every 100,000 to 1,000,000 per generation is a mutant that resists the gef gene’s action. Will the clones win or will the mutants win? Biotechnologists have helped push the odds in favor of “good” clones over “bad” mutants by inserting two gef genes into P. putida, which lengthens the odds of resistance to one cell in every 100,000,000.
Anthrax
If E. coli is the world’s most bioengineered bacterium, Bacillus anthracis is the most feared because it causes the disease anthrax. B. anthracis joins various viruses, parasites, other bacteria, and toxins (made by bacteria or fungi) on a list of potential bioterrorism threats. Not only does the B. anthracis toxin cause lethal effects in humans, but the bacterium’s ability to form endospores helps it out-survive other pathogens. Endospore-formation keeps the cells alive and yet resistant to chemicals, irradiation, and antibiotics.
B. anthracis begins as a laboratory culture like other bacteria. To make endospores, a microbiologist stresses the cells by heating the culture broth. The cells begin forming endospores within minutes of this stress. The microbiologist can freeze-dry the spores to make a brown to off-white powder; the color depends on the type of medium that had been used for growing the cells.
The dry, odorless, and lethal powder has caused significant concern in the United States, especially since anthrax was used as a presumed weapon distributed in mail in 2001. Security teams at airports and public buildings now search for unidentified powders as possible anthrax.
As a bioweapon, other pathogens work better than anthrax. The pathogen causes illness if it enters the body through a wound in the skin, by ingestion, or inhalation, with inhalation being the likely route of infection for a bioweapon. The skin route would be impractical for a terrorist and putting anthrax into food or water becomes ineffective because of a phenomenon called the dilution effect. Community water supplies and food products contain such large volumes that a terrorist would find it impossible to contaminate either with a dose big enough to kill. The endospores require a large dose to cause infection in people, so food and especially water would dilute them to harmless levels. A terrorist would furthermore be hard-pressed to perform the laborious culturing and freeze-drying steps needed to make a significant amount of endospores.
Disease by inhalation has caused greater concern because it has already been shown to cause most anthrax cases, the contamination of postal letters in 2001, for instance. But not everyone who gets infected develops disease. People who do get sick cannot transmit it to others because anthrax is noncontagious. Even though B. anthracis grows easily in a laboratory, all other characteristics of this microbe make it a poor bioweapon. Therefore, the most feared bacterium is not as big a threat to a large population of people as many believe.
Why we will always need bacteria
White biotechnology offers the greatest hope of integrating bacteria into industry in a way to positively affect the environment. The use of bacteria to perform activities now carried out by strong acids and organic solvents will drastically cut down on the chemical wastes flowing into rivers, soils, and groundwaters. Many industrial steps must take place at several hundred degrees, which consumes large amounts of energy. Bacteria substitute biodegradable enzymes for caustic chemicals and work at mild temperatures, and they do it quietly. Microbial fermentations also produce heat that can be rerouted into the manufacturing facility to reduce overall energy use.
Bacteria are white biotech’s raw material. Rather than watch truck or trainloads of chemicals roll toward manufacturing plants, neighbors of a white biotech company might spot a person carrying a single vial of freeze-dried bacteria. From that point, the bacteria regenerate themselves. In fact, ancient societies might wonder why present-day industry bothers with their noxious mix of materials and wastes. Bacteria already make almost every compound humans find important, even plastic. Pseudomonas species make polyesters called polyhydroxyalkanoates (PHAs) from sugars or lipids found in nature. The bacteria use the large compounds as a storage form of carbon and energy and as the sticky binder in biofilm.
Industrial interest in PHAs increases or decreases with oil prices because oil serves as a cheap precursor for making most plastics. As oil prices rise, PHAs become more attractive for making soft containers such as shampoo bottles. But PHA production is not inexpensive due to the costs of nutrients for growing the bacteria and methods for harvesting the polymer.
Bacillus megaterium and Alcaligenes eutrophus lead a group of diverse species that produce nature’s most abundant PHA, polyhydroxybutyrate (PHB). Bacteria excrete higher amounts of PHB under stress, probably as a protective coating around the cells. The narrow environmental conditions that induce the bacteria to turn on their PHB genes make this a very expensive natural product compared with plastics derived from fossil fuels. PHBs are compatible with human tissue because they do not cause allergic reactions, and they are pliable. These attributes make PHBs good choices for medical equipment such as flexible tubing and intravenous bags. To reach this promising future for biodegradable plastics, white biotechnology will be called upon to find the secrets of bacterial metabolism that lead to the cost-effective production of PHAs.
Some manufacturing processes have changed little since the dawn of the Industrial Revolution. Of all aspects of society, manufacturing lags the furthest behind in converting traditional processes into more sustainable methods. For this important change to take place the most self-sufficient organisms on Earth might well lead the way.
