This is the most controversial section in the book so far. Genetic algorithms are based on the biological theory of evolution (see http://en.wikipedia.org/wiki/Evolutionary_algorithm). This type of algorithm is useful for searching and optimization. For instance, we can use it to find the optimal parameters for a regression or classification problem.
Humans and other life forms on Earth carry genetic information in chromosomes. Chromosomes are frequently modeled as strings. A similar representation is used in genetic algorithms. The first step is to initialize the population with random individuals and related representation of genetic information. We can also initialize with already-known candidate solutions for the problem. After that, we go through many iterations called generations. During each generation, individuals are selected for mating based on a predefined fitness function. The fitness function evaluates how close an individual is to the desired solution.
Two genetic operators generate new genetic information:
- Crossover: This occurs via mating and creates new children. We will explain one-point crossover here. This process takes a piece of genetic information from one parent and a complementary piece from the other parent. For example, if the information is represented by 100 list elements, crossover may take the first 80 element of the first parent and the last 20 from the other parent. It is possible in genetic algorithms to produce children from more than two parents. This is an area under research (refer to Eiben, A. E. et al. Genetic algorithms with multi-parent recombination, Proceedings of the International Conference on Evolutionary Computation – PPSN III. The Third Conference on Parallel Problem Solving from Nature: 78–87. ISBN 3-540-58484-6, 1994).
- Mutation: This is controlled by a fixed mutation rate. This concept is explained in several Hollywood movies and popular culture. Mutation is rare and often detrimental or even fatal. However, sometimes mutants can acquire desirable traits. In certain cases, the trait can be passed on to future generations.
Eventually, the new individuals replace the old population and we can start a new iteration. In this example, we will use the Python DEAP library. Install DEAP as follows:
$ sudo pip install deap $ pip freeze|grep deap deap==1.0.1
Start by defining a Fitness subclass that maximizes fitness:
creator.create("FitnessMax", base.Fitness, weights=(1.0,))Then, define a template for each individual in the population:
creator.create("Individual", array.array, typecode='d', fitness=creator.FitnessMax)DEAP has the concept of a toolbox, which is a registry of necessary functions. Create a toolbox and register the initialization functions, as follows:
toolbox = base.Toolbox()
toolbox.register("attr_float", random.random)
toolbox.register("individual", tools.initRepeat, creator.Individual, toolbox.attr_float, 200)
toolbox.register("populate", tools.initRepeat, list, toolbox.individual)The first function generates floating-point numbers between 0 and 1. The second function creates an individual with a list of 200 floating point numbers. The third function creates a list of individuals. This list represents the population of possible solutions for a search or optimization problem.
In a society, we want "normal" individuals, but also people like Einstein. In Chapter 3, Statistics and Linear Algebra, we were introduced to the shapiro() function, which performs a normality test. For an individual to be normal, we require that the normality test p-value of his or her list to be as high as possible. The following code defines the fitness function:
def eval(individual):
return shapiro(individual)[1],Let's define the genetic operators:
toolbox.register("evaluate", eval)
toolbox.register("mate", tools.cxTwoPoint)
toolbox.register("mutate", tools.mutFlipBit, indpb=0.1)
toolbox.register("select", tools.selTournament, tournsize=4)The following list will give you an explanation about the preceding genetic operators:
evaluate: This operator measures the fitness of each individual. In this example, the p-value of a normality test is used as a fitness score.mate: This operator produces children. In this example, it uses two-point crossover.mutate: This operator changes an individual at random. For a list of Boolean values, this means that some values are flipped fromTruetoFalseand vice versa.select: This operator selects the individuals that are allowed to mate.
In the preceding code snippet, we specified that we are going to use two-point crossover and the probability of an attribute to be flipped. Generate 400 individuals as the initial population:
pop = toolbox.populate(n=400)
Now start the evolution process, as follows:
hof = tools.HallOfFame(1)
stats = tools.Statistics(key=lambda ind: ind.fitness.values)
stats.register("max", np.max)
algorithms.eaSimple(pop, toolbox, cxpb=0.5, mutpb=0.2, ngen=80, stats=stats, halloffame=hof)The program reports statistics including the maximum fitness for each generation. We specified the crossover probability, mutation rate, and maximum generations after which to stop. The following is an extract of the displayed statistics report:
gen nevals max 0 400 0.000484774 1 245 0.000776807 2 248 0.00135569 … 79 250 0.99826 80 248 0.99826
As you can see, we start out with distributions that are far from normal, but eventually we get an individual with the following histogram:
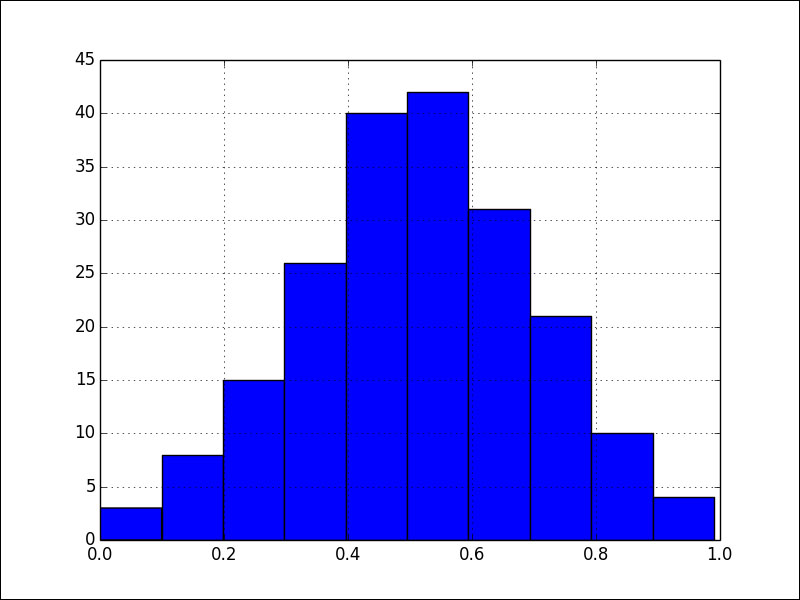
Refer to the gen_algo.py file in this book's code bundle:
import array
import random
import numpy as np
from deap import algorithms
from deap import base
from deap import creator
from deap import tools
from scipy.stats import shapiro
import matplotlib.pyplot as plt
creator.create("FitnessMax", base.Fitness, weights=(1.0,))
creator.create("Individual", array.array, typecode='d', fitness=creator.FitnessMax)
toolbox = base.Toolbox()
toolbox.register("attr_float", random.random)
toolbox.register("individual", tools.initRepeat, creator.Individual, toolbox.attr_float, 200)
toolbox.register("populate", tools.initRepeat, list, toolbox.individual)
def eval(individual):
return shapiro(individual)[1],
toolbox.register("evaluate", eval)
toolbox.register("mate", tools.cxTwoPoint)
toolbox.register("mutate", tools.mutFlipBit, indpb=0.1)
toolbox.register("select", tools.selTournament, tournsize=4)
random.seed(42)
pop = toolbox.populate(n=400)
hof = tools.HallOfFame(1)
stats = tools.Statistics(key=lambda ind: ind.fitness.values)
stats.register("max", np.max)
algorithms.eaSimple(pop, toolbox, cxpb=0.5, mutpb=0.2, ngen=80, stats=stats, halloffame=hof)
print shapiro(hof[0])[1]
plt.hist(hof[0])
plt.grid(True)
plt.show()