 K-means with H2O
by Alberto Boschetti, Luca Massaron, Bastiaan Sjardin, John Hearty, Prateek Joshi
Python: Real World Machine Learning
K-means with H2O
by Alberto Boschetti, Luca Massaron, Bastiaan Sjardin, John Hearty, Prateek Joshi
Python: Real World Machine Learning
- Python: Real World Machine Learning
- Table of Contents
- Python: Real World Machine Learning
- Python: Real World Machine Learning
- Credits
- Preface
- I. Module 1
- 1. The Realm of Supervised Learning
- Introduction
- Preprocessing data using different techniques
- Label encoding
- Building a linear regressor
- Computing regression accuracy
- Achieving model persistence
- Building a ridge regressor
- Building a polynomial regressor
- Estimating housing prices
- Computing the relative importance of features
- Estimating bicycle demand distribution
- 2. Constructing a Classifier
- Introduction
- Building a simple classifier
- Building a logistic regression classifier
- Building a Naive Bayes classifier
- Splitting the dataset for training and testing
- Evaluating the accuracy using cross-validation
- Visualizing the confusion matrix
- Extracting the performance report
- Evaluating cars based on their characteristics
- Extracting validation curves
- Extracting learning curves
- Estimating the income bracket
- 3. Predictive Modeling
- 4. Clustering with Unsupervised Learning
- Introduction
- Clustering data using the k-means algorithm
- Compressing an image using vector quantization
- Building a Mean Shift clustering model
- Grouping data using agglomerative clustering
- Evaluating the performance of clustering algorithms
- Automatically estimating the number of clusters using DBSCAN algorithm
- Finding patterns in stock market data
- Building a customer segmentation model
- 5. Building Recommendation Engines
- Introduction
- Building function compositions for data processing
- Building machine learning pipelines
- Finding the nearest neighbors
- Constructing a k-nearest neighbors classifier
- Constructing a k-nearest neighbors regressor
- Computing the Euclidean distance score
- Computing the Pearson correlation score
- Finding similar users in the dataset
- Generating movie recommendations
- 6. Analyzing Text Data
- Introduction
- Preprocessing data using tokenization
- Stemming text data
- Converting text to its base form using lemmatization
- Dividing text using chunking
- Building a bag-of-words model
- Building a text classifier
- Identifying the gender
- Analyzing the sentiment of a sentence
- Identifying patterns in text using topic modeling
- 7. Speech Recognition
- 8. Dissecting Time Series and Sequential Data
- Introduction
- Transforming data into the time series format
- Slicing time series data
- Operating on time series data
- Extracting statistics from time series data
- Building Hidden Markov Models for sequential data
- Building Conditional Random Fields for sequential text data
- Analyzing stock market data using Hidden Markov Models
- 9. Image Content Analysis
- Introduction
- Operating on images using OpenCV-Python
- Detecting edges
- Histogram equalization
- Detecting corners
- Detecting SIFT feature points
- Building a Star feature detector
- Creating features using visual codebook and vector quantization
- Training an image classifier using Extremely Random Forests
- Building an object recognizer
- 10. Biometric Face Recognition
- Introduction
- Capturing and processing video from a webcam
- Building a face detector using Haar cascades
- Building eye and nose detectors
- Performing Principal Components Analysis
- Performing Kernel Principal Components Analysis
- Performing blind source separation
- Building a face recognizer using Local Binary Patterns Histogram
- 11. Deep Neural Networks
- Introduction
- Building a perceptron
- Building a single layer neural network
- Building a deep neural network
- Creating a vector quantizer
- Building a recurrent neural network for sequential data analysis
- Visualizing the characters in an optical character recognition database
- Building an optical character recognizer using neural networks
- 12. Visualizing Data
- 1. The Realm of Supervised Learning
- II. Module 2
- 1. Unsupervised Machine Learning
- 2. Deep Belief Networks
- 3. Stacked Denoising Autoencoders
- 4. Convolutional Neural Networks
- 5. Semi-Supervised Learning
- 6. Text Feature Engineering
- 7. Feature Engineering Part II
- 8. Ensemble Methods
- 9. Additional Python Machine Learning Tools
- A. Chapter Code Requirements
- III. Module 3
- 1. First Steps to Scalability
- 2. Scalable Learning in Scikit-learn
- 3. Fast SVM Implementations
- 4. Neural Networks and Deep Learning
- The neural network architecture
- Neural networks and regularization
- Neural networks and hyperparameter optimization
- Neural networks and decision boundaries
- Deep learning at scale with H2O
- Deep learning and unsupervised pretraining
- Deep learning with theanets
- Autoencoders and unsupervised learning
- Summary
- 5. Deep Learning with TensorFlow
- 6. Classification and Regression Trees at Scale
- 7. Unsupervised Learning at Scale
- 8. Distributed Environments – Hadoop and Spark
- 9. Practical Machine Learning with Spark
- A. Introduction to GPUs and Theano
- A. Bibliography
- Index
Here, we're comparing the K-means implementation of H2O with Scikit-learn. More specifically, we will run the mini-batch experiment using H2OKMeansEstimator, the object for K-means available in H2O. The setup is similar to the one shown in the PCA with H2O section, and the experiment is the same as seen in the preceding section:
In:import h2o
from h2o.estimators.kmeans import H2OKMeansEstimator
h2o.init(max_mem_size_GB=4)
def testH2O_kmeans(X, k):
temp_file = tempfile.NamedTemporaryFile().name
np.savetxt(temp_file, np.c_[X], delimiter=",")
cls = H2OKMeansEstimator(k=k, standardize=True)
blobdata = h2o.import_file(temp_file)
tik = time.time()
cls.train(x=range(blobdata.ncol), training_frame=blobdata)
fit_time = time.time() - tik
os.remove(temp_file)
return fit_time
piece_of_dataset = pd.read_csv(census_csv_file, iterator=True).get_chunk(500000).drop('caseid', axis=1).as_matrix()
time_results = {4: [], 8:[], 12:[]}
dataset_sizes = [20000, 200000, 500000]
for dataset_size in dataset_sizes:
print "Dataset size:", dataset_size
X = piece_of_dataset[:dataset_size,:]
for K in [4, 8, 12]:
print "K:", K
fit_time = testH2O_kmeans(X, K)
time_results[K].append(fit_time)
plt.plot(dataset_sizes, time_results[4], 'r', label='K=4')
plt.plot(dataset_sizes, time_results[8], 'g', label='K=8')
plt.plot(dataset_sizes, time_results[12], 'b', label='K=12')
plt.xlabel("Training set size")
plt.ylabel("Training time")
plt.legend(loc=0)
plt.show()
testH2O_kmeans(100000, 100)
h2o.shutdown(prompt=False)
Out: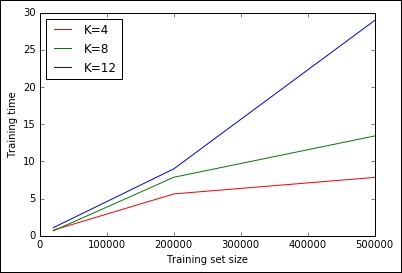
Thanks to the H2O architecture, its implementation of K-means is very fast and scalable and able to perform the clustering of the 500K point datasets in less than 30 seconds for all the selected Ks.
-
No Comment
..................Content has been hidden....................
You can't read the all page of ebook, please click here login for view all page.